Visualization in ChimeraX/ArtiaX
Contents
Visualization in ChimeraX/ArtiaX¶
nextPYP produces all the necessary files to visualize a refined map into the original tomogram positions using ChimeraX/ArtiaX.
Step 1: Download the necessary files¶
Select the refinement block where you will extract the particle locations from. Click on the Edit menu for that block and select the Show filesystem location to find out the location of the project in the filesystem.
From this location, you will need to download the following files to your local computer:
Tomogram reconstruction (
mrc/tile_series_name.rec)Refined map (
frealign/maps/block_name_r{class_number}_{iteration_number}_crop.mrc)Corresponding particle orientations (
frealign/artiax/tile_series_name_K1.star)
Step 2: Load data into ChimeraX/ArtiaX¶
Open ChimeraX (we assume the ArtiaX plugin is already installed)
Open the tomogram file
tilt_series_name.rec- Run the following commands in the ChimeraX shell:
volume permuteAxes #1 xzyvolume flip #2 axis z(this step is only necessary when the dataset has virions)
Go to the ArtiaX tab and
Launchthe pluginIn the Tomograms section (main ArtiaX panel on the left), select model #3 (permuted z flip) from the
Add Modeldropdown menu and clickAdd!Go to to the ArtiaX options panel on the right, and set the
Pixel Sizefor the Current Tomogram to the binned pixel size (10.8 for the EMPIAR-10164 tutorial) and clickApplyFrom the Particles List section (main ArtiaX panel on the left), select
Open List ...and browse to the location of the.starfileGo to the ArtiaX options panel on the right, select the Select/Manipulate tab and set the
Originof thePixelsize Factorsto the unbinned pixel size of the data (1.35 for the EMPIAR-10164 tutorial)From the Geometric Models section (main ArtiaX panel on the left), go to
Open Geomodel ...and select the refined mapGo to the ArtiaX options panel on the right, in the Visualization tab, set the
Use Modelfrom the dropdown menu to the refined map and clickAttach ModelFrom the Color Settings section, select
Colormapand choose the attributerlnLogLikelihoodContributionfrom the dropdown menu
If everything went well, you should obtain a result similar to this:
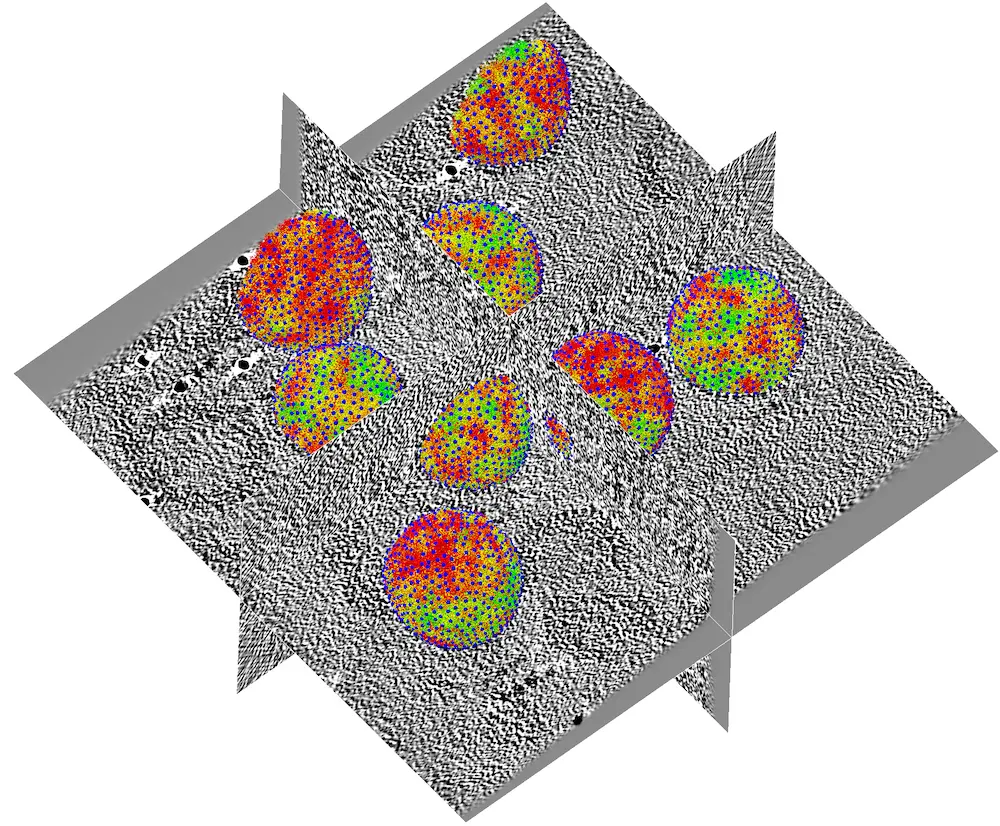
Tip
Depending on the dimensions of the refined map and the number of particles in the tomogram, you may need to downsample the map to make ChimeraX more responsive.